S-NITROSATION
In its classical signaling role, NO is captured by the heme cofactor of soluble guanylate cyclase (sGC), activating sGC to produce the secondary messenger cyclic GMP (cGMP). However, mounting evidence points toward an alternative, cGMP-independent NO signaling pathway in which the S-nitrosation of cysteine residues regulates protein structure and activity. S-Nitrosation has been implicated in a broad spectrum of diseases, including cancer, diabetes, and other cardiovascular, pulmonary, and neurological disorders, yet the mechanism by which nitrosothiols are formed in vivo is unknown. In vitro, non-specific cysteine nitrosation occurs readily. In vivo nitrosation, however, is far more selective for specific cysteines and is driven by factors beyond thiol reactivity. We postulate that nitrosation selectivity is driven by protein-protein or protein-small molecule interactions that align a nitrosothiol with a free thiol for transnitrosation reactions.
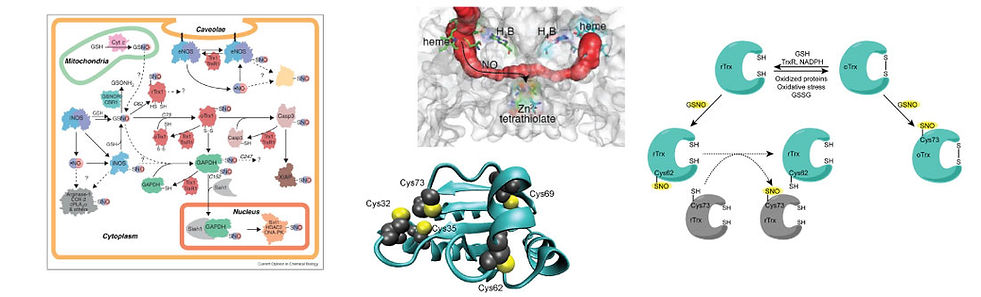
Transnitrosation reactions require an “initiating” nitrosothiol. Nitric oxide synthases (NOSs) are potential candidates for the initial formation of nitrosothiols as all three mammalian NOS isoforms selectively form nitrosothiols at their Zn2+-tetrathiolate cysteines. We recently developed a kinetic model of NOS S-nitrosation. In this model, NO synthesized at the heme cofactor is partitioned between release into solution and NOS auto-S-nitrosation. The results suggested that NOS S-nitrosation is both a mechanism to control NOS activity and to generate physiological nitrosothiols.
The inducible NOS isoform (iNOS) has been shown to participate in protein–protein interaction-mediated S-nitrosation reactions with cyclooxygenase-2 (COX-2) and arginase-1. Furthermore, procaspase-3 and iNOS participate in an NO-dependent protein–protein interaction. As caspase-3 is known to be nitrosated on its active-site cysteine, iNOS might directly transnitrosate caspase-3. We are broadly interested in discovering and characterizing novel targets of NOS transnitrosation.

Recent genome sequencing revealed that many bacterial species also possess NOS-like proteins. While most of these bacterial NOS proteins contain only a single domain with high sequence homology to the oxygenase domain of mammalian NOS, a handful of recently-discovered bacterial NOS proteins are fused to reductase domains. We are generally interested in these novel bacterial NOS proteins. Do these bacterial NOS proteins produce NO? What substrates and cofactors are required for their activity? How do the mechanisms of NO synthesis compare and contrast to the mammalian NOS enzymes? What role do these bacterial NOS proteins play in signaling?

Together, these studies will provide compelling evidence for S-nitrosation as a signaling-competent, reversible post-translational modification driven by selective protein-protein and protein-small molecule interactions, and will set the stage for a molecular understanding of S-nitrosation in vivo.
Publications:
Bruegger JJ, Smith BC, Wynia-Smith SL, Marletta, MA. Comparative and integrative metabolomics reveal that S-nitrosation inhibits physiologically relevant metabolic enzymes. JBC 2018, 293, 6282-6296.
Smith BC, Marletta MA. Mechanisms of S-nitrosothiol formation and selectivity in nitric oxide signaling. Curr Opin Chem Biol 2012, 16:498-506.
Smith BC, Fernhoff, NB, Marletta MA. Mechanisms and kinetics of nitric oxide synthase auto-S-nitrosation and inactivation. Biochemistry 2012, 51:1028-40.
Barglow KT, Knutson CG, Wishnok JS, Tannenbaum SR, Marletta MA. Site-specific and redox-controlled S-nitrosation of thioredoxin. Proc Natl Acad Sci USA 2011, 108:E600-6.
Mitchell DA, Michel T, Marletta MA. Effects of S-nitrosation of nitric oxide synthase. Adv Exp Biol 2007, 1:151-79.
Mitchell DA, Morton SU, Fernhoff NB, Marletta MA. Thioredoxin is required for S-nitrosation of procaspase-3 and the inhibition of apoptosis in Jurkat cells. Proc Natl Acad Sci USA 2007, 104:11609-14.
Mitchell DA, Morton SU, Marletta MA. Design and characterization of an active site selective caspase-3 transnitrosating agent. ACS Chem Biol 2006, 1:659-65.
Erwin PA, Mitchell DA, Sartoretto J, Marletta MA, Michel T. Subcellular targeting and differential S-nitrosylation of endothelial nitric-oxide synthase. J Biol Chem 2006, 281:151-7.
Mitchell DA, Marletta MA. Thioredoxin catalyzes the S-nitrosation of the caspase-3 active site cysteine. Nat Chem Biol 2005, 1:154-8.
Mitchell DA, Erwin PA, Michel T, Marletta MA. S-Nitrosation and regulation of inducible nitric oxide synthase. Biochemistry 2005, 44:4636-47.